As I said 6 months ago, 2019 is a tough year to write a blog, because this was going to be a complex regional election year and therefore a time of political promises, hence tenure offers too. Now the preliminary offers have been made, elections have passed, but the timing has slightly shifted toward 2020. So I may have the time, but not really any benefit of dedicating too much effort to the blog, and a lot of potential benefit of dedicating any time to evaluable scientific work.
On the other hand, I saw some potential benefit for publishing texts with ISBNs, hence the updates to the text and the preparation of these printed copies of the books, just in case. While Spain’s accreditation agency has some hard rules for becoming a tenured professor, especially for medical associates (whose years of professional experience are almost worthless compared to published peer-reviewed papers), it is quite flexible in assessing one’s merits.
However, regional and/or autonomous entities are not, and need an official identifier and preferably printed versions to evaluate publications, such as an ISBN for books. I took thus some time about a month ago to update the texts and supplementary materials, to publish a printed copy of the books with Amazon. The first copies have arrived, and they look good.
Corrections and Additions
Titles
I have changed the names and order of the books, as I intended for the first publication – as some of you may have noticed when the linguistic book was referred to as the third volume in some parts. In the first concept I just wanted to emphasize that the linguistic work had priority over the rest. Now the whole series and the linguistic volume don’t share the same name, and I hope this added clarity is for the better, despite the linguistic volume being the third one.
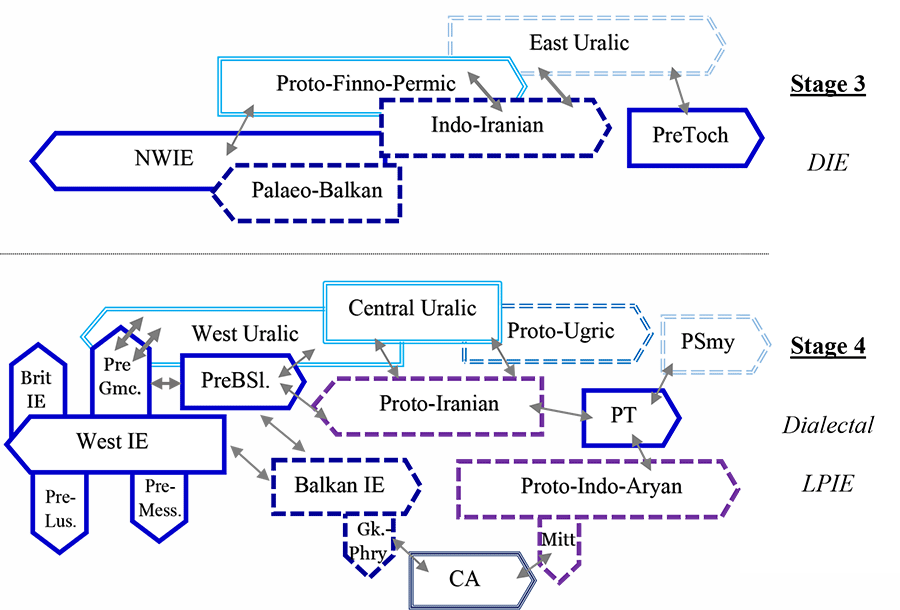
Uralic dialects
I have changed the nomenclature for Uralic dialects, as I said recently. I haven’t really modified anything deeper than that, because – unlike adding new information from population genomics – this would require for me to do a thorough research of the most recent publications of Uralic comparative grammar, and I just can’t begin with that right now.
Anyway, the use of terms like Finno-Ugric or Finno-Samic is as correct now for the reconstructed forms as it was before the change in nomenclature.
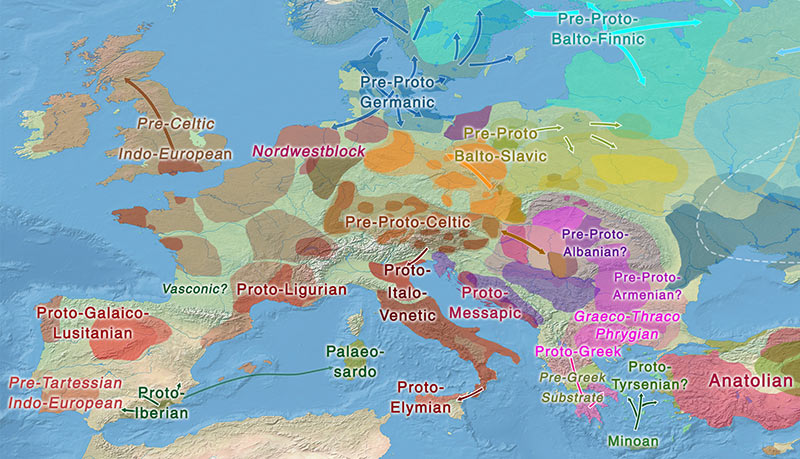
Mediterranean
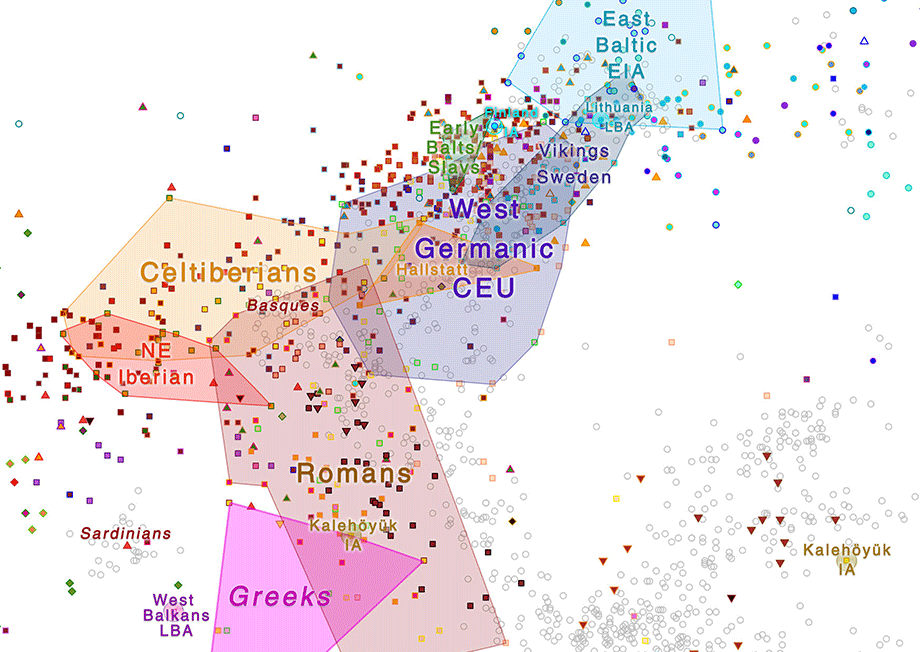
The most interesting recent genetic data has come from Iberia and the Mediterranean. Lacking direct data from the Italian Peninsula (and thus from the emergence of the Etruscan and Rhaetian ethnolinguistic community), it is becoming clearer how some quite early waves of Indo-Europeans and non-Indo-Europeans expanded and shrank – at least in West Iberia, West Mediterranean, and France.
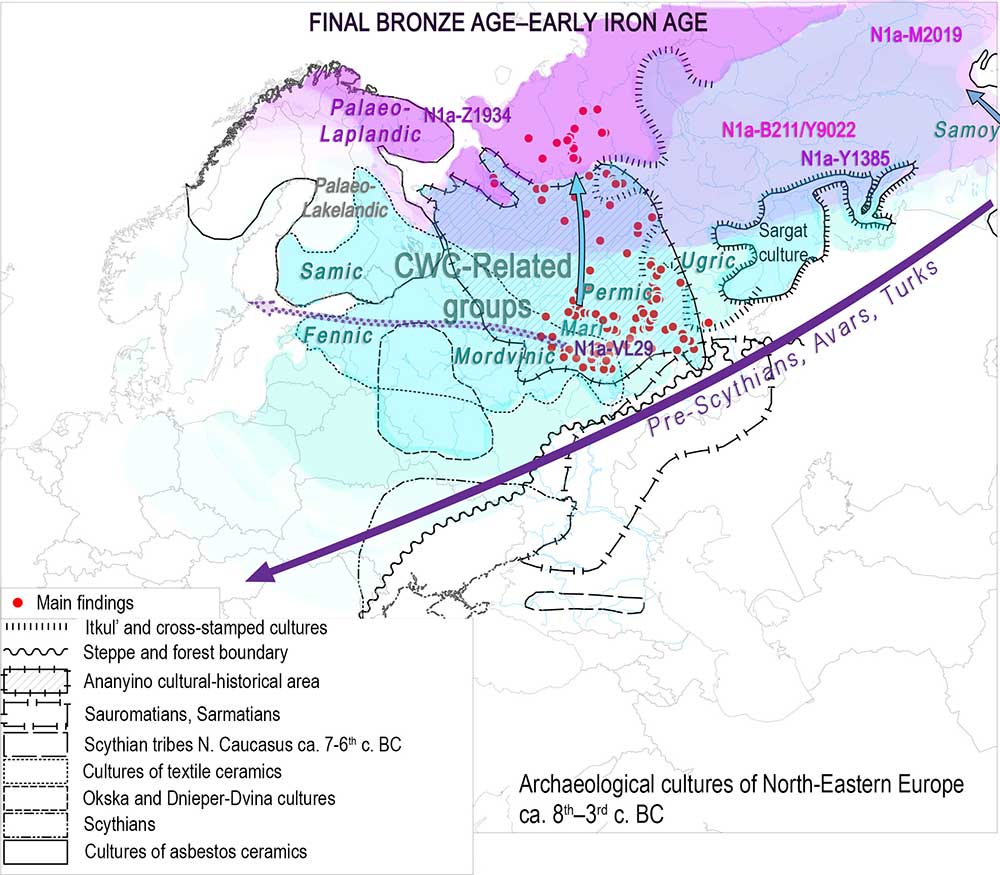
Finno-Ugric
Some of the main updates to the text have been made to the sections on Finno-Ugric populations, because some interesting new genetic data (especially Y-DNA) have been published in the past months. This is especially true for Baltic Finns and for Ugric populations.
Balto-Slavic
Consequently, and somehow unsurprisingly, the Balto-Slavic section has been affected by this; e.g. by the identification of Early Slavs likely with central-eastern populations dominated by (at least some subclades of) hg. I2a-L621 and E1b-V13.
Maps
I have updated some cultural borders in the prehistoric maps, and the maps with Y-DNA and mtDNA. I have also added one new version of the Early Bronze age map, to better reflect the most likely location of Indo-European languages in the Early European Bronze Age.
As those in software programming will understand, major changes in the files that are used for maps and graphics come with an increasing risk of additional errors, so I would not be surprised if some major ones would be found (I already spotted three of them). Feel free to communicate these errors in any way you see fit.
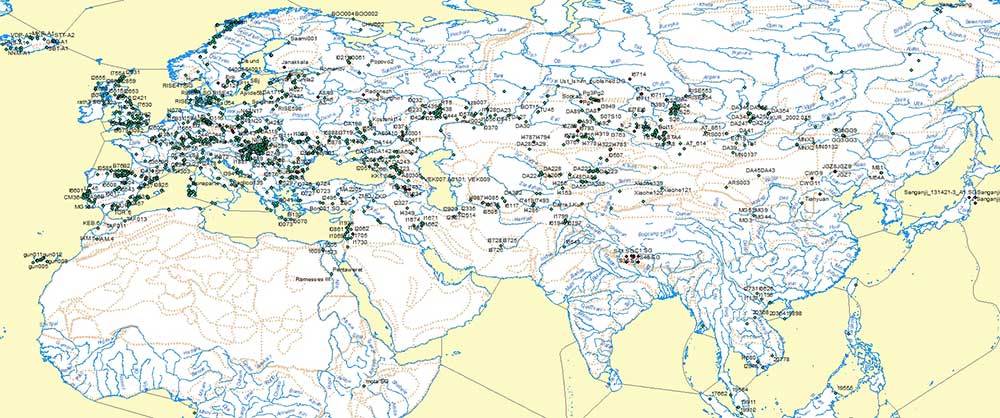
SNPs
I have selected more conservative SNPs in certain controversial cases.
I have also deleted most SNP-related footnotes and replaced them with the marking of each individual tentative SNP, leaving only those footnotes that give important specific information, because:
- My way of referencing tentative SNP authors did not make it clear which samples were tentative, if there were more than one.
- It was probably not necessary to see four names repeated 100 times over.
- Often I don’t really know if the person I have listed as author of the SNP call is the true author – unless I saw the full SNP data posted directly – or just someone who reposted the results.
- Sometimes there are more than one author of SNPs for a certain sample, but I might have added just one for all.
For a centralized file to host the names of those responsible for the unofficial/tentative SNPs used in the text – and to correct them if necessary -, readers will be eventually able to use Phylogeographer‘s tool for ancient Y-DNA, for which they use (partly) the same data I compiled, adding Y-Full‘s nomenclature and references. You can see another map tool in ArcGIS.
NOTE. As I say in the text, if the final working map tool does not deliver the names, I will publish another supplementary table to the text, listing all tentative SNPs with their respective author(s).
If you are interested in ancient Y-DNA and you want to help develop comprehensive and precise maps of ancient Y-DNA and mtDNA haplogroups, you can contact Hunter Provyn at Phylogeographer.com. You can also find more about phylogeography projects at Iain McDonald’s website.
Graphics
I have also added more samples to both the “Asian” and the “European” PCAs, and to the ADMIXTURE analyses, too.
I previously used certain samples prepared by amateurs from BAM files (like Botai, Okunevo, or Hittites), and the results were obviously less than satisfactory – hence my criticism of the lack of publication of prepared files by the most famous labs, especially the Copenhagen group.
Fortunately for all of us, most published datasets are free, so we don’t have to reinvent the wheel. I criticized genetic labs for not releasing all data, so now it is time for praise, at least for one of them: thank you to all responsible at the Reich Lab for this great merged dataset, which includes samples from other labs.
NOTE. I would like to make my tiny contribution here, for beginners interested in working with these files, so I will update – whenever I have time – the “How To” sections of this blog for PCAs, PCA3d, and ADMIXTURE.
ADMIXTURE
For unsupervised ADMIXTURE in the maps, a K=5 is selected based on the CV, giving a kind of visual WHG : NWAN : CHG/IN : EHG : ENA, but with Steppe ancestry “in between”. Higher K gave worse CV, which I guess depends on the many ancient and modern samples selected (and on the fact that many samples are repeated from different sources in my files, because I did not have time to filter them all individually).
I found some interesting component shared by Central European populations in K=7 to K=9 (from CEU Bell Beakers to Denmark LN to Hungarian EBA to Iberia BA, in a sort of “CEU BBC ancestry” potentially related to North-West Indo-Europeans), but still, I prefer to go for a theoretically more correct visualization instead of cherry-picking the ‘best-looking’ results.
Since I made fun of the search for “Siberian ancestry” in coloured components in Tambets et al. 2018, I have to be consistent and preferred to avoid doing the same here…
qpAdm
In the first publication (in January) and subsequent minor revisions until March, I trusted analyses and ancestry estimates reported by amateurs in 2018, which I used for the text adding my own interpretations. Most of them have been refuted in papers from 2019, as you probably know if you have followed this blog (see very recent examples here, here, or here), compelling me to delete or change them again, and again, and again. I don’t have experience from previous years, although the current pattern must have been evidently repeated many times over, or else we would be still talking about such previous analyses as being confirmed today…
I wanted to be one step ahead of peer-reviewed publications in the books, but I prefer now to go for something safe in the book series, rather than having one potentially interesting prediction – which may or may not be right – and ten huge mistakes that I would have helped to endlessly redistribute among my readers (online and now in print) based on some cherry-picked pairwise comparisons. This is especially true when predictions of “Steppe“- and/or “Siberian“-related ancestry have been published, which, for some reason, seem to go horribly wrong most of the time.
I am sure whole books can be written about why and how this happened (and how this is going to keep happening), based on psychology and sociology, but the reasons are irrelevant, and that would be a futile effort; like writing books about glottochronology and its intermittent popularity due to misunderstood scientist trends. The most efficient way to deal with this problem is to avoid such information altogether, because – as you can see in the current revised text – they wouldn’t really add anything essential to the content of these books, anyway.
Continue reading
Official site of the book series:
A Song of Sheep and Horses: eurafrasia nostratica, eurasia indouralica