Open access Genetic Landscape of Slovenians: Past Admixture and Natural Selection Pattern, by Maisano Delser et al. Front. Genet. (2018).
Interesting excerpts (emphasis mine):
Samples
Overall, 96 samples ranging from Slovenian littoral to Lower Styria were genotyped for 713,599 markers using the OmniExpress 24-V1 BeadChips (Figure 1), genetic data were obtained from Esko et al. (2013). After removing related individuals, 92 samples were left. The Slovenian dataset has been subsequently merged with the Human Origin dataset (Lazaridis et al., 2016) for a total of 2163 individuals.
Y chromosome
First, Y chromosome genetic diversity was assessed. A total of 52 Y chromosomes were analyzed for 195 SNPs. The majority of individuals (25, 48.1%) belong to the haplogroup R1a1a1a (R-M417) while the second major haplogroup is represented by R1b (R-M343) including 15 individuals (28.8%). Twelve samples are assigned to haplogroup I (I M170): five and two samples belong to haplogroup I2a (I L460) and I1 (I M253), respectively, while the remaining five samples did not have enough information to be further assigned.
PCA
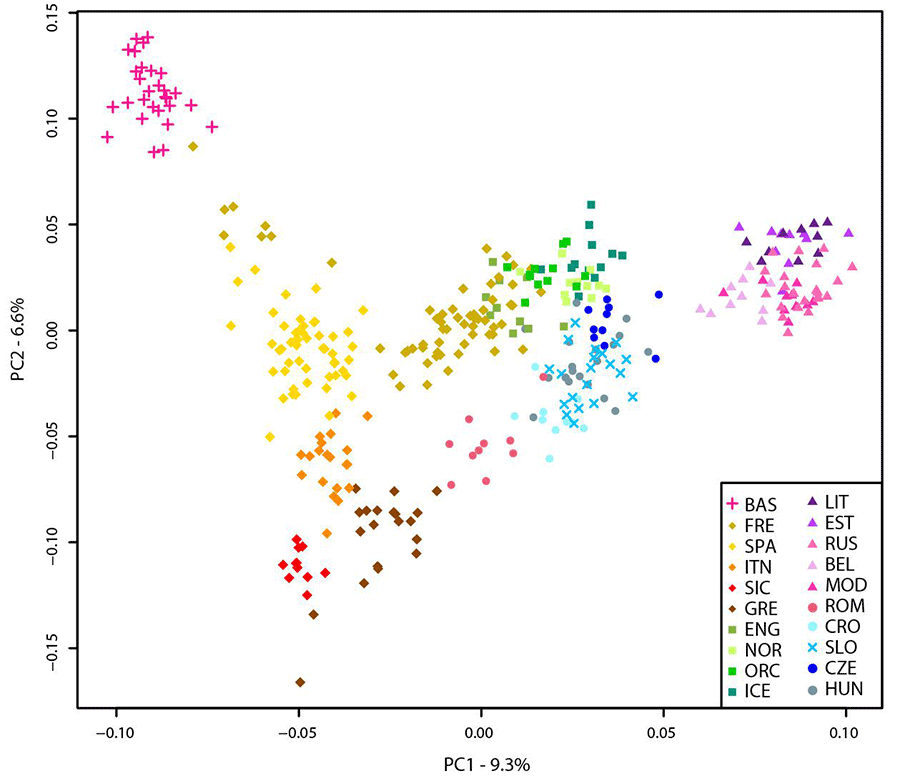
Considering the unbalanced sample size of the Slovenian population compared to the other populations included in the dataset, a subset of 20 Slovenian individuals randomly sampled was used.
All Slovenian samples group together with Hungarians, Czechs, and some Croatians (“Central-Eastern European” cluster) as also suggested by the PCA. All Basque individuals with few French and Spanish cluster together (“Basque” cluster) while a “Northern-European” cluster is made of the majority of French, English, Icelanders, Norwegians, and Orcadians. Five populations contributed to the “Eastern-European” cluster including Belarusians, Estonians, Lithuanians, Mordovians, and Russians. Western and South Europe is split into two cluster: the first (“Western European” cluster) includes all Spanish individuals, few French, and some Italians (North Italy) while the second (“Southern-European” cluster) groups Sicilians, Greeks, some Croatians, Romanians, and some Italians (North Italy).
Admixture Pattern and Migration
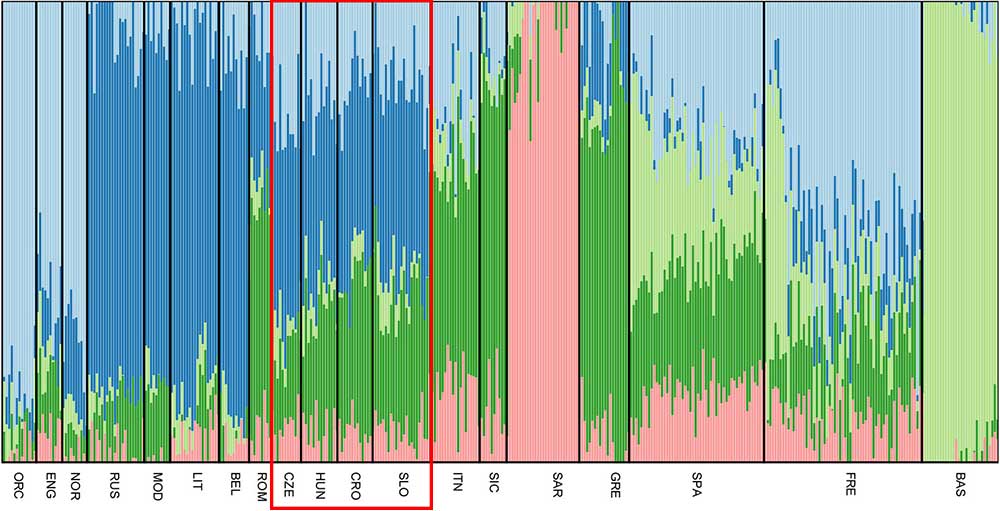
All Slovenian individuals share common pattern of genetic ancestry, as revealed by ADMIXTURE analysis. The three major ancestry components are the North East and North West European ones (light blue and dark blue, respectively, Figure 3), followed by a South European one (dark green, Figure 3). Contribution from the Sardinians and Basque are present in negligible amount. The admixture pattern of Slovenians mimics the one suggested by the neighboring Eastern European populations, but it is different from the pattern suggested by North Italian populations even though they are geographically close.
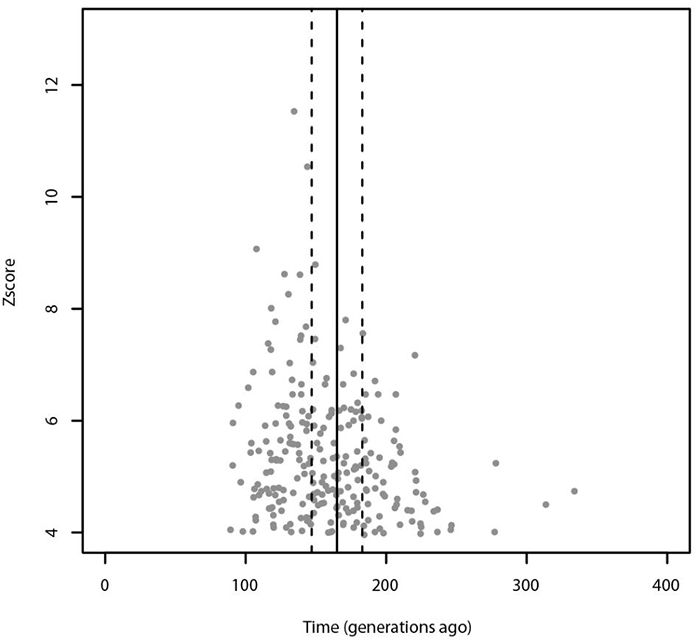
Using ALDER, the most significant admixture event was obtained with Russians and Sardinians as source populations and it happened 135 ± 9.31 generations ago (Z-score = 11.54). (…) When tested for multiple admixture events (MALDER), we obtained evidence for one admixture event 165.391 ± 17.1918 generations ago corresponding to ∼2620 BCE (CI: 3101–2139) considering a generation time of 28 years (Figure 4), with Kalmyk and Sardinians as sources.
We then modeled the Slovenian population as target of admixture of ancient individuals from Haak et al. (2015) while computing the f3(Ancient 1, Ancient 2, Slovenian) statistic. The most significant signal was obtained with Yamnaya and HungaryGamba_EN (Z-score = -10.66), followed by MA1 with LBK_EN (Z-score -9.7) and Yamnaya with Stuttgart (Z-score = -8.6) used as possible source populations (Supplementary Figure 5).
We found a significant signal of admixture by using both pairs as ancient sources. Specifically, for the pair Yamnaya and Hungary_EN the admixture event is dated at 134.38 ± 23.69 generations ago (Z-score = 5.26, p-value of 1.5e-07) while for Yamnaya and LBK_EN at 153.65 ± 22.19 generations ago (Z-score = 6.92, p-value 4.4e-12). Outgroup f3 with Yamnaya put Slovenian population close to Hungarians, Czechs, and English, indicating a similar shared drift between these population with the Steppe populations (Supplementary Figure 6).
Not that any of this would come as a surprise, but:
- R1a-M458 and some R1a-Z280 (xR1a-Z92) lineages (found among Slovenes) were associated with the Slavic expansion, likely with the Prague-Korchak culture, originally stemming probably from peoples of the Lusatian culture. Other R1a-Z280 lineages remained associated with Uralic peoples, and some became Slavicized only recently.
- PCA keeps supporting the common cluster of certain West, South, and East Slavs in a “Central-Eastern European” cluster, distinct from the “North-Eastern European” cluster formed by modern Finno-Ugrians, as well as ancient Finno-Ugrians of north-eastern Europe who were only recently Slavicized.
- Admixture supports the same ancient ‘western’ (a core West+South+East Slavic) cluster, and the admixture event with Yamna + Hungary_EN is logically a proxy for Yamna Hungary being at the core of ancestral Central-East population movements related to Bell Beakers in the mid- to late 3rd millennium.
The theory that East Slavs are at the core of the Slavic expansion makes no sense, in terms of archaeology (see Florin Curta’s dismissal of those recent eastern ‘Slavic’ finds, his commentary on 19th century Pan-Slavic crap, or his book on Slavic migrations), in terms of ancient DNA (the earliest Slavs sampled cluster with modern West Slavs, distant from the steppe cluster, unlike Finno-Ugrians), or in terms of modern DNA.
I don’t know where exactly this impulse for the theory of Russia being the cradle of Slavs comes from today (although there are some obvious political trends to revive 19th c. ideas), but it was always clear for everyone, including Russians, that East Slavs had migrated to the east and north and assimilated indigenous Finno-Ugrians, apart from Turkic-, Iranian-, and Caucasian-speaking peoples to the east. Genetics is only confirming what was clear from other disciplines long ago.
Related
- On the origin and spread of haplogroup R1a-Z645 from eastern Europe
- Consequences of O&M 2018 (III): The Balto-Slavic conundrum in Linguistics, Archaeology, and Genetics
- Olalde et al. and Mathieson et al. (Nature 2018): R1b-L23 dominates Bell Beaker and Yamna, R1a-M417 resurges in East-Central Europe during the Bronze Age
- The concept of “Outlier” in Human Ancestry (III): Late Neolithic samples from the Baltic region and origins of the Corded Ware culture
- The significance of the Tollense Valley in Bronze Age North-East Germany
- The Tollense Valley battlefield: the North European ‘Trojan war’ that hints to western Balto-Slavic origins
- The Indo-European demic diffusion model, and the “R1b – Indo-European” association
- Our monograph on North-West Indo-European (first draft) is out
- Early Indo-Iranian formed mainly by R1b-Z2103 and R1a-Z93, Corded Ware out of Late PIE-speaking migrations
- Y-DNA haplogroup R1b-Z2103 in Proto-Indo-Iranians?